
When using phylogenies in spatial conservation prioritisation, we need to link the phylogeny with distribution data. Increasingly, distribution data is used to predict where species occur across the landscape using a species distribution model (SDM). SDMs are currently underused in conservation, but have great potential for a variety of applications from threatened species management to conservation planning. Our recent paper shows how to use SDMs with a phylogeny in spatial conservation planning (this method could also be used for a variety of applications linking phylogenies and SDMs).
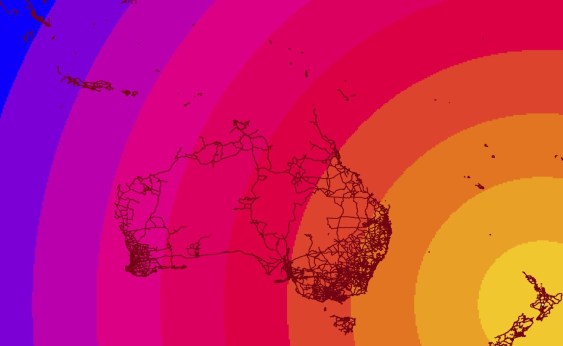
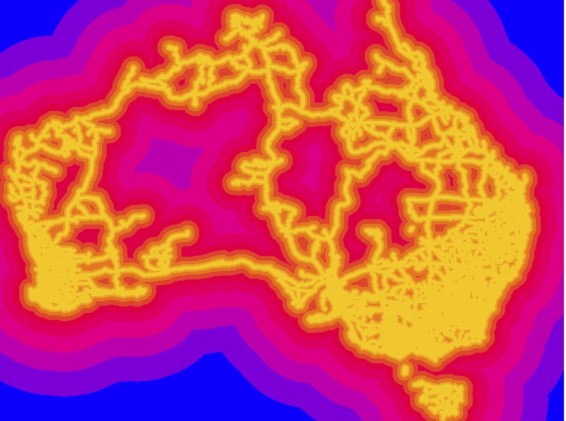
An SDM models the response of a species to a set of predictor variables (usually environmental variables). The model can be extended across a landscape with a probability of occurrence of species in grid cells**. The external branches (tips) of the phylogeny correspond to a particular taxon (let’s assume we have a species-level tree). Therefore, each external branch can simply be the probability of that species occurring in each cell (a,b,c,e,f in figure above). Now, for the internal branches. Continue reading Linking species distribution models (SDM) and phylogenies

 We have a new
We have a new 
