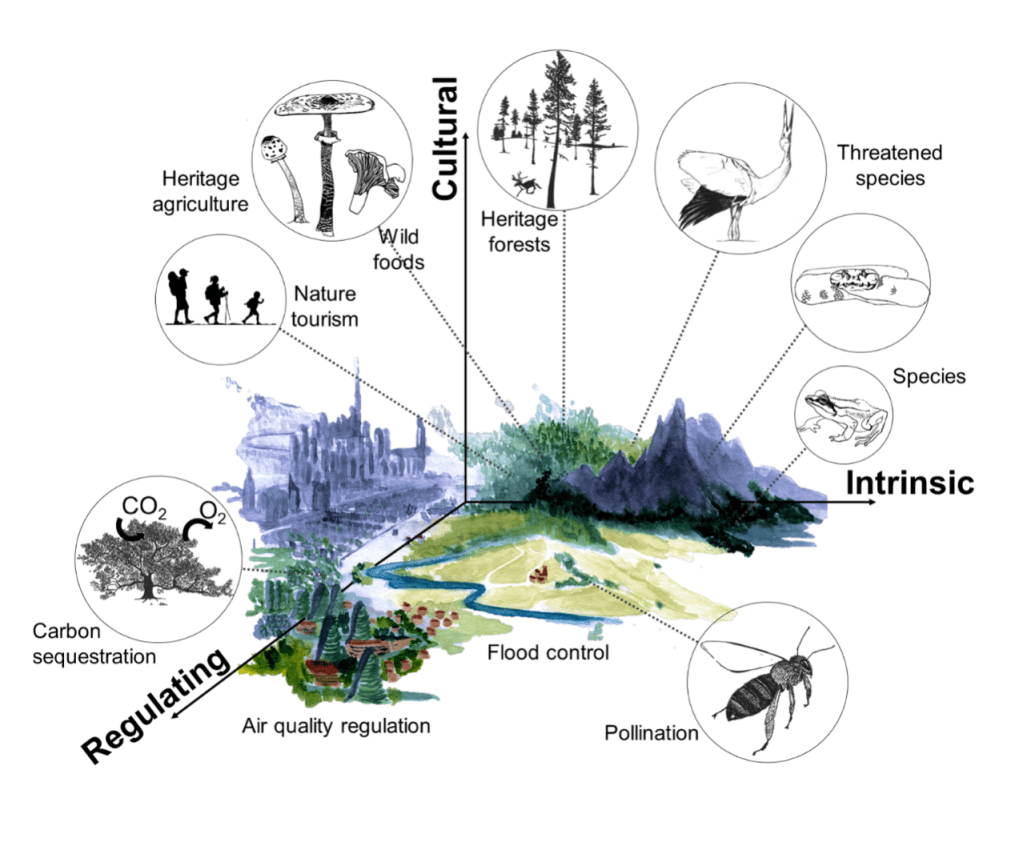
The many values of nature: Intrinsic value reflects the high value attributed to biodiversity (e.g. species) and ecosystem functioning, independently of human presence. Cultural value reflects the high value placed on natural heritage and traditions rooted in nature (e.g. the diversity of mushroom and plant species for gathering, heritage landscapes, nature tourism). Regulating value places high value on ecosystems that contribute to natural processes that sustain societies, such as flood control, air quality regulation, pollination, and climate regulation. (Illustrated by Dr. Camille Martinez-Almoyna)
Many countries have committed to ’30 by 30’- the goal of protecting 30% of land by 2030, but what exactly we aim to protect is still debated. Two major goals are to protect biodiversity for the sake of biodiversity (nature for nature) or to protect the biodiversity that directly benefits humans (nature for people), but few studies actually compare how these contrasting viewpoints change our priorities. Are the places in need of protection similar for biodiversity and nature’s contributions to people (NCP)?
We assembled large datasets on biodiversity (all terrestrial animals in Europe) and a set of cultural and regulating NCP across Europe, and compared the spatial overlap of resulting priority areas.
Specifically, we looked at: (i) biodiversity, represented by 785 terrestrial vertebrate species, including 124 threatened species; (ii) regulating NCP, represented by carbon sequestration, air quality regulation, flood prevention and pollination (Fig. S4); and (iii) cultural NCP, represented by heritage forests, heritage agriculture, foraging for wild foods (mushrooms and plants), and nature-based tourism.
We found that priorities are very different for biodiversity compared to NCP, and many NCP also differ from each other. Only a few small ecosystems are priorities for both (mainly in Mediterranean forests). Importantly, we also found that identifying priorities for biodiversity captures NCP much better than the reverse. If we only protect NCP, many important species would be lost.
We also identified priority areas given what is already protected by the Natura 2000 network–so candidates for expanding protection. Natura 2000 is one of the densest conservation networks in the world, covering about 20% of the EU, but it only protects 30% of the key areas for NCP, and 36% of vertebrate species distributions on average. Our results show that an additional 5% put into protection could double the current protection of vertebrate species and regulation NCP, and would protect almost 75% of threatened species distributions on average (Figure below).
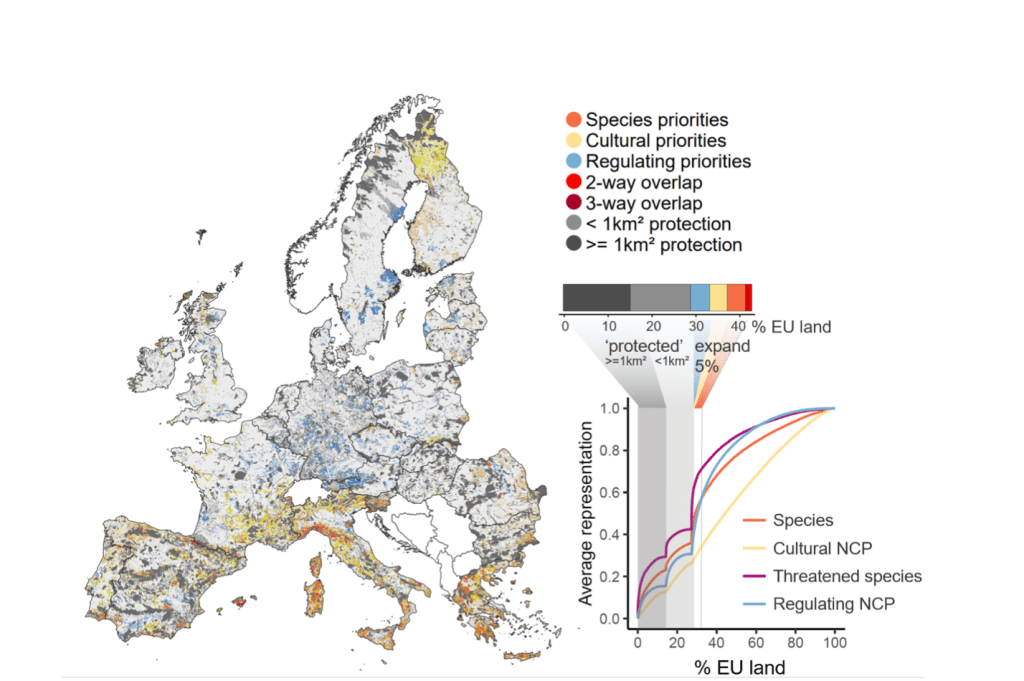
Priority areas to improve the protection of vertebrate species and NCP in Europe. On the left, the coloured areas show currently unprotected sites that are essential for different nature values (orange, all species; yellow, cultural; blue, regulation). Areas of overlap between different nature values are in red. The existing Natura 2000 areas are in grey. On the right, the bar chart quantifies the area of the EU occupied by each type of priority, and areas of overlap, outside Natura 2000 areas. At the bottom right, the performance curves quantify the average representation for each value of nature as area is added, taking into account Natura 2000 areas (in grey).
The key areas identified have the potential to improve the conservation of species and NCP on a continental scale. These ecosystems are potentially home to a many terrestrial vertebrates and provide essential ecosystem services. They are irreplaceable (i.e. they contain species that are not found elsewhere) and they optimise the protection of each species and NCP in a minimum of area. Our study is a preliminary step towards a better integration of nature’s values in conservation. The designation of protected areas will depend on multiple stakeholders and on socio-economic feasibility – dimensions that we did not explore in our study, but which need to be considered. Furthermore, international coordination for nature conservation will be vital to halt the decline of global biodiversity
Contribution to Sustainable development goals
- SDG 13: The priority areas we have identified are relevant for climate change mitigation and adaptation, as they aim to protect (among other things):
- terrestrial ecosystems that sequester CO2
- ecosystems that can control floods, which is relevant to climate change adaptation because extreme climate events, such as heavy rainfall, are likely to become increasingly frequent and unpredictable.
- SDG 15: We identify ecosystems that are essential for the survival of terrestrial vertebrate species. Protecting these areas may prevent extinctions of terrestrial vertebrate species.
- SDG 17: Our results confirm the importance of international coordination to conserve species and NCP at the European scale.

 The world’s biodiversity is in crisis. Species are declining at an alarming rate. And this is happening at just the time we are really beginning to understand this diversity through an unprecedented cataloging and compiling of information. Data repositories are filled with hundreds of thousands of entries about species, where they live, how they live, and who they are related to. And this is only the beginning. New DNA-based surveys are exploding onto the scene and our ideas and understanding of biodiversity are improving everyday.
The world’s biodiversity is in crisis. Species are declining at an alarming rate. And this is happening at just the time we are really beginning to understand this diversity through an unprecedented cataloging and compiling of information. Data repositories are filled with hundreds of thousands of entries about species, where they live, how they live, and who they are related to. And this is only the beginning. New DNA-based surveys are exploding onto the scene and our ideas and understanding of biodiversity are improving everyday.
 We have a new
We have a new 